Disease strainsMutationsIt is quite possible that a virus changes its nature during the course of an epidemic. Usually such random mutations lead nowhere and the novel strain dies out quickly, but occasionally a mutation produces a strain that is more efficient, i.e. transmits with higher probability. Such a mutation asks the whole stability question in yet a different way, because a situation that is stable for the original disease may not be for the novel strain.Using a seeding, it is possible to introduce a more agressive strain into the simulation and subsequently even tag its effect. SeedingsCreate the following config file as the baseline epidemic:
This creates a relatively slow-spreading disease which is from timestep 400 on contained by a low-level but long measure. If nothing else happens, the epidemic stops over time without spreading across the whole grid. Note the transmission_factor keyword - this is a different way to specify the effect of a measure. While transmission sets the transmission probability to a fixed value (regardless of the strain causing the infection), transmission_factor multiplies the value of the strain, i.e. it treats original and novel strain of the disease differently. Which of these options is true in nature? It depends on how the transmission probability is reduced by the measure. In a situation where contacts are reduced, but people behave normally during a contact, it does not matter which strain is in the situation - most contacts lead to an infection, the measure just makes this sufficiently rare. If on the other hand the transmission probability is based on reducing the amount of virus exposure by e.g. protective equipment, limiting exposure time or keeping safe distances, then a more agressive strain would react less to the measure. In the following, we assume the latter possibility. Using
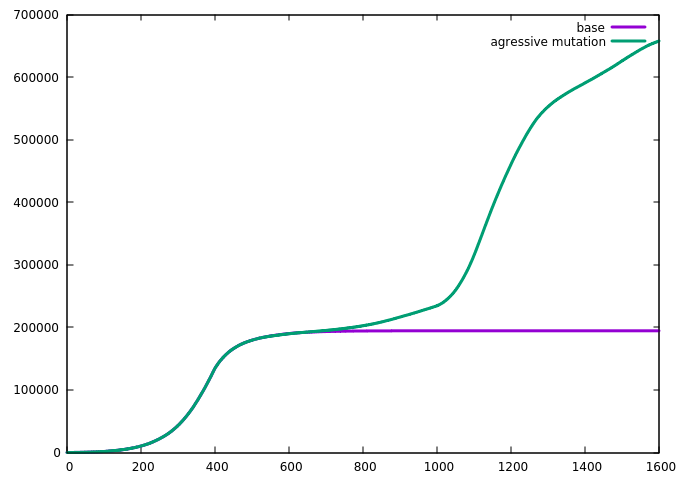
we introduce a faster-spreading virus strain into the simulation while the containing measure is active. The parameters are chosen such that while the original strain can not spread under the measure, the novel strain can. The result is as follows:
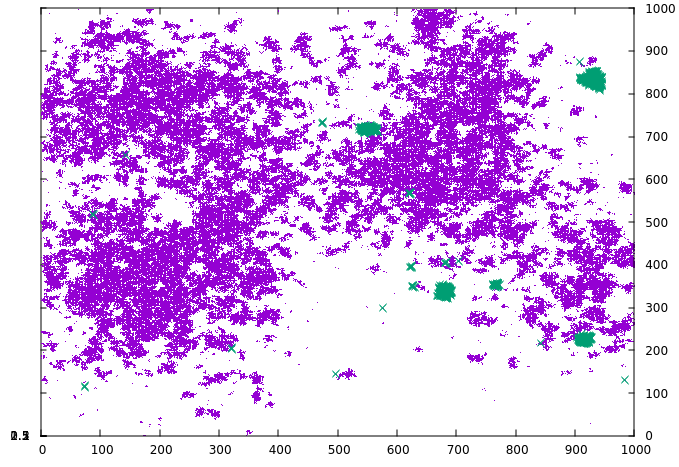
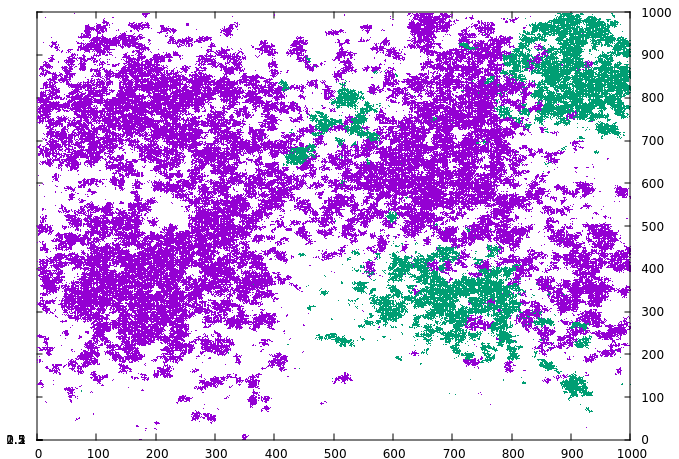
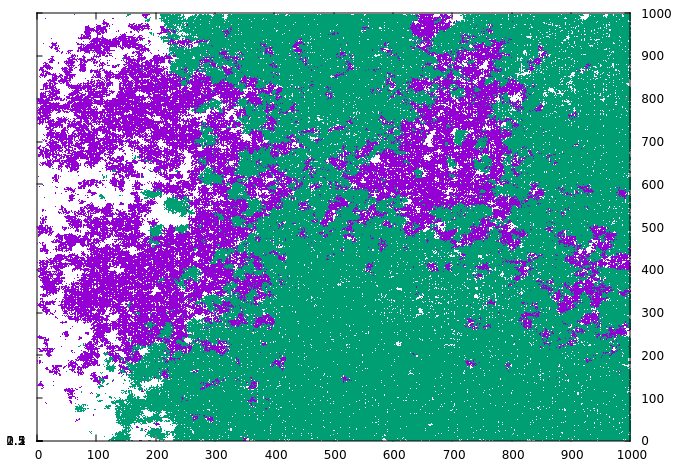
The novel strain manages to spread under the containment measure and then quickly grows when the measure ends in timestep 1000. By using the tag_index keyword, we can visualize this on the grid. If the index is defined, every infected grid cell writes the specified index rather than a one when a data file is written, and this can be utilized by the plot software to distinguish. As can be seen in the following series, the novel strain (green) manages to grow in small pockets under the containment measure while the original strain (violet) is largely static:
By the time the measure ends and the original strain has stopped growing, the novel strain has already grown substantially.
Finally, the novel strain has spread across most of the grid, except for the regions where the original strain was particularly active and has created protective pockets of immunity.
RemarksIt should be obvious that any stability analysis can be rendered obsolete by the appearance of a sufficiently agressive novel strain. Moreover, the simulation so far has assumed that being infected by either the original or the novel strain grants immunity against both. This need not be the case in nature however, as the influenza virus amply demonstrates, immunity may only be partial - or not there at all.
Continue with Vaccination campaigns. Back to main index Back to science Back to numerical epidemic Created by Thorsten Renk 2017-2021 - see the disclaimer, privacy statement and contact information. |